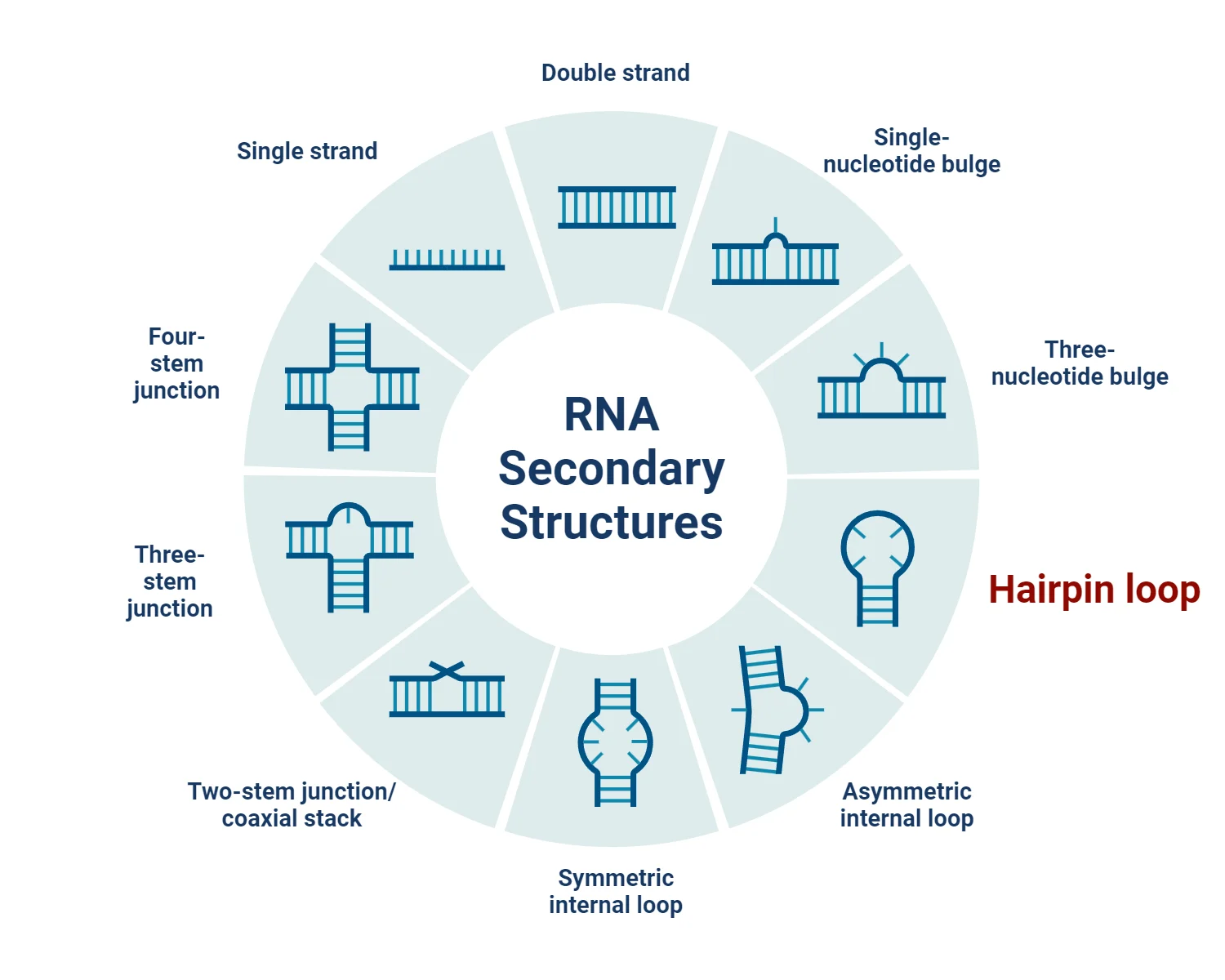
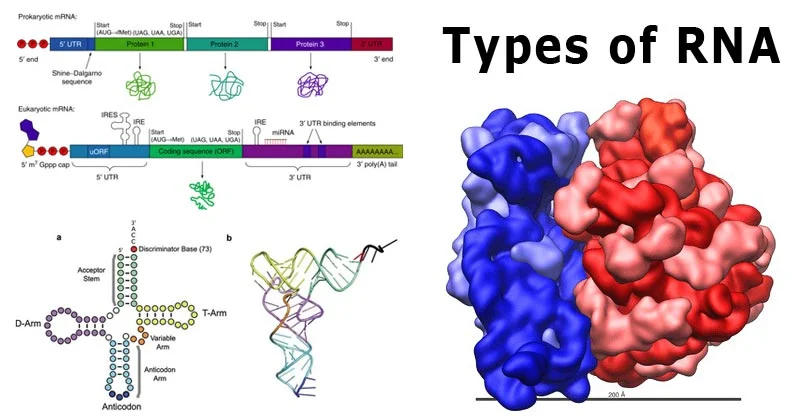
Stem-loop, also known as a hairpin loop, is a common secondary structure found in nucleic acids, particularly in single-stranded RNA and DNA molecules. This structure plays a critical role in various biological processes such as gene regulation, RNA folding, and the functioning of certain ribozymes. The formation of stem-loop structures results from the intramolecular base pairing within a single-stranded molecule, which forms a stem (paired bases) and a loop (unpaired bases) that resembles a hairpin. This article delves into the properties, types, examples, and uses of stem-loop structures in nucleic acids.
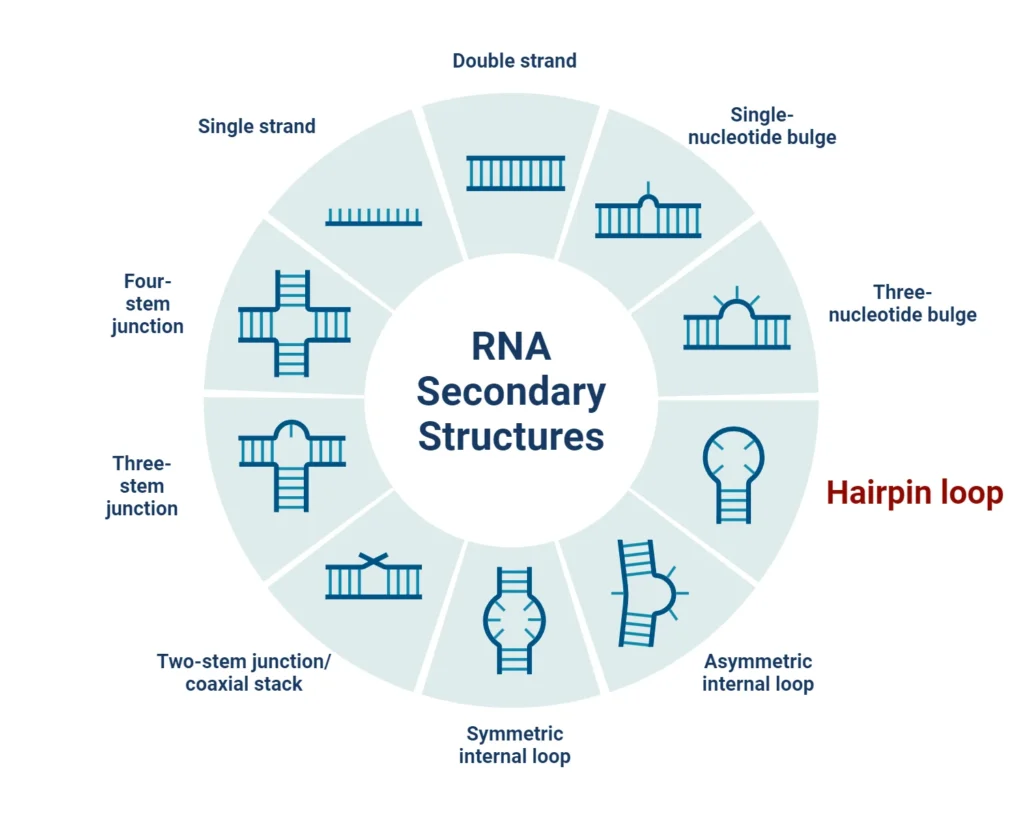
What is a Stem-loop (Hairpin Loop)?
A stem-loop structure refers to a specific pattern formed by the intramolecular base pairing of nucleotides, typically within palindromic sequences. In this structure:
- The stem is composed of paired nucleotide bases, forming a double helix.
- The loop is an unpaired region that ends the stem, creating a hairpin-like structure.
- This structure can occur in both RNA and DNA, though it is more commonly found in RNA.
Stem-loops are integral to the secondary structure of many RNA molecules, affecting their stability, function, and interactions with proteins or other RNAs.
Characteristics of Stem-loop (Hairpin Loop)
- Form in Single-stranded Nucleic Acids:
- Stem-loops are primarily observed in single-stranded RNA, though they can also form in single-stranded DNA.
- Composed of Two Parts:
- Stem: A base-paired region, typically forming a double helix with complementary base-pairing (A-U, G-C).
- Loop: A region of unpaired bases that creates a loop-like structure at the end of the stem.
- Base-pairing and Stability:
- The stem forms complementary base pairs between nucleotides within the same strand.
- The loop often contains 7 to 20 unpaired bases, and the sequence of these bases can influence the stability of the hairpin structure.
- Variability in Loop Length:
- The loop length can vary and is typically between 4 to 8 bases for optimal stability.
- Short loops (less than 4 bases) are generally unstable, while large loops can lead to destabilization.
- Types of Hairpin Loops:
- Depending on the number of unpaired bases in the loop, stem-loop structures may be referred to as tri-, tetra-, penta-loops, etc.
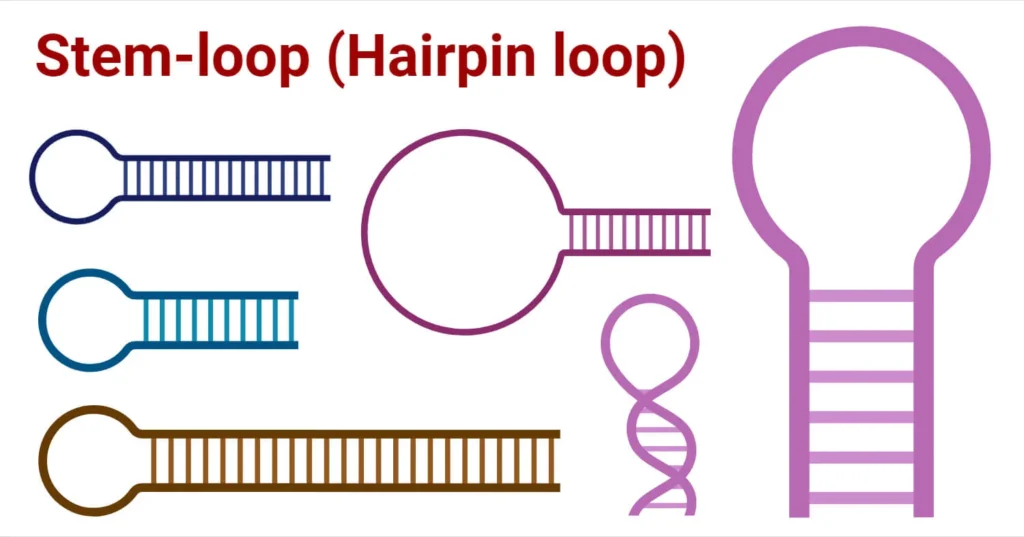
Origin of RNA Hairpin
RNA hairpin structures can arise through two primary mechanisms:
- Transcription by RNA Polymerase:
- RNA polymerase transcribes inverted repeat DNA sequences, leading to RNA folding into a hairpin loop structure.
- RNA-Dependent RNA Polymerase:
- In certain viral replication processes, RNA serves as a template for RNA-dependent RNA polymerase, which synthesizes the second stem strand, leading to the formation of long, perfect double-stranded RNA hairpins.
These mechanisms allow RNA molecules to adopt complex structures that are essential for gene regulation, replication, and other cellular processes.
Molecules with Loop Structures
Stem-loop structures are found in a variety of RNA and DNA molecules. Notable examples include:
- tRNA (Transfer RNA):
- tRNA molecules contain three hairpin loops that resemble a three-leaf clover.
- One of these loops contains the anticodon, which plays a crucial role in recognizing and decoding mRNA codons during protein synthesis.
- RNA Pseudoknots:
- RNA pseudoknots are secondary structures that feature nested stem-loop formations.
- These structures play roles in translation and replication.
- Ribozymes:
- Certain ribozymes, such as the hammerhead ribozyme, rely on stem-loop structures for their catalytic activities.
- Ribozymes often contain multiple stem-loop regions that are critical for self-cleavage.
- Prokaryotic 5′ UTRs (Untranslated Regions):
- In prokaryotes, hairpin loops in 5′ UTRs are important for regulating translation initiation and interacting with regulatory proteins.
Stability Factors of Stem-loop (Hairpin Loop)
The stability of stem-loop structures depends on several factors:
- Base Pairing:
- Guanine-cytosine (G-C) pairings are more stable than adenine-uracil (A-U) pairings, contributing to the overall stability of the stem.
- Sequence Composition:
- Specific base sequences within the stem can influence stability. For example, certain sequences like the UUCG tetraloop are particularly stable.
- Loop Length:
- The optimal loop length for stability is generally between 4 to 8 bases. Short loops (less than 4 bases) are generally unstable, while larger loops can cause the structure to destabilize.
- Base Stacking:
- Base stacking interactions between adjacent base pairs within the stem help stabilize the helix structure.
- Temperature Sensitivity:
- The stability of RNA hairpins can also depend on temperature. Higher temperatures may cause the hairpin structure to unfold, disrupting the base pairs and loop structure.
Examples of Stem-loop (Hairpin Loop)
- Prokaryotic Transcription Termination:
- In prokaryotes, hairpin loops at the end of certain transcriptional regions can cause RNA polymerase to dissociate from the DNA, terminating transcription.
- tRNA Structure:
- tRNA contains three distinct hairpin loops. The anticodon loop is particularly important in decoding mRNA during translation. The other two loops help stabilize the overall tRNA structure.
- DNA Hairpin:
- Example of a DNA hairpin sequence:
- Palindromic sequence:
CCTGCXXXXXXXGCAGG - Forms the following hairpin structure:
- C G
C G
T A
G C
C G
X X
X X
X X
X
- C G
- Palindromic sequence:
- Example of a DNA hairpin sequence:
- RNA Sequence Example:
- Example of a longer RNA sequence forming a hairpin loop:
GCCGCGGGCCGAAAAAACCCCCCCGGCCCGCGGC
- Example of a longer RNA sequence forming a hairpin loop:
Significance of Stem-loop (Hairpin Loop)
Significance of RNA Stem-loop Structures:
- Gene Regulation:
- RNA stem-loop structures regulate gene expression both in cis (within the same RNA molecule) and in trans (affecting other molecules).
- Biosensor and Diagnostic Tools:
- Stem-loop structures can be used in biosensors. Well-designed oligonucleotides with stem-loop structures can open in the presence of complementary sequences, allowing detection of nucleic acids or proteins without the need for labeling.
- Translation Regulation:
- Short RNA hairpins in mRNA are involved in translation regulation by modulating mRNA localization, translation initiation, and stability.
- RNA Folding and Nucleation Sites:
- Hairpin loops are crucial for RNA folding, influencing RNA-protein recognition and the formation of functional RNA structures.
Significance of DNA Hairpins:
- Single-stranded Phages:
- In single-stranded phages, DNA hairpins are used during replication, helping the phage to replicate its genome efficiently.
- Cruciform DNA:
- DNA regions with hairpin motifs, known as cruciform DNA, are involved in processes such as genome translocation and replication initiation.
- Diagnostic and Biotechnological Tools:
- DNA hairpin probes are widely used in diagnostics and biotechnology due to their ability to discriminate between matched and mismatched DNA sequences.
Uses of Stem-loop Structures
- Diagnostics:
- Hairpin loop structures are widely used in biosensors for the detection of nucleic acids and proteins, utilizing their ability to open in the presence of complementary sequences.
- Gene Regulation:
- Stem-loop structures play a pivotal role in regulating mRNA stability, localization, and translation.
- Biotechnology and Molecular Biology:
- Hairpin DNA probes are essential in biotechnology for detecting specific target sequences.
- Stem-loop structures are used to design specific oligonucleotides in diagnostic assays, ensuring high specificity in sequence detection.
- Therapeutic Applications:
- Stem-loop structures are increasingly being explored in therapeutic applications, including gene silencing, and in RNA-based vaccines and therapeutics.
Stem-loop (hairpin loop) structures are crucial elements in the molecular biology of RNA and DNA. These structures not only aid in the regulation of gene expression but also serve as powerful tools in diagnostic and biotechnological applications. Their ability to stabilize nucleic acid secondary structures and participate in RNA folding makes them indispensable in various biological processes.